At a Hay Festival event sponsored by the Royal Society, Director of External Affairs Roger Highfield interviewed the President of the Society, Nobelist Venki Ramakrishnan.
The ribosome is a complex biological machine that sits at the nexus of history, information, life and health. It is a molecular factory that has toiled in organisms since the dawn of life billions of years ago, one that plays a central role in all living things and one that, according to Venki Ramakrishnan, could provide the key to combating the global scourge of antibiotic resistance.
Venki is a leading light among those who have been unravelling the secrets of this vast chemical machine. His team at the Medical Research Council’s Laboratory of Molecular Biology in Cambridge has uncovered key details of this protein-making factory, life’s oldest and most universal assembly of molecules, paving the way for his 2009 Nobel Prize in chemistry.
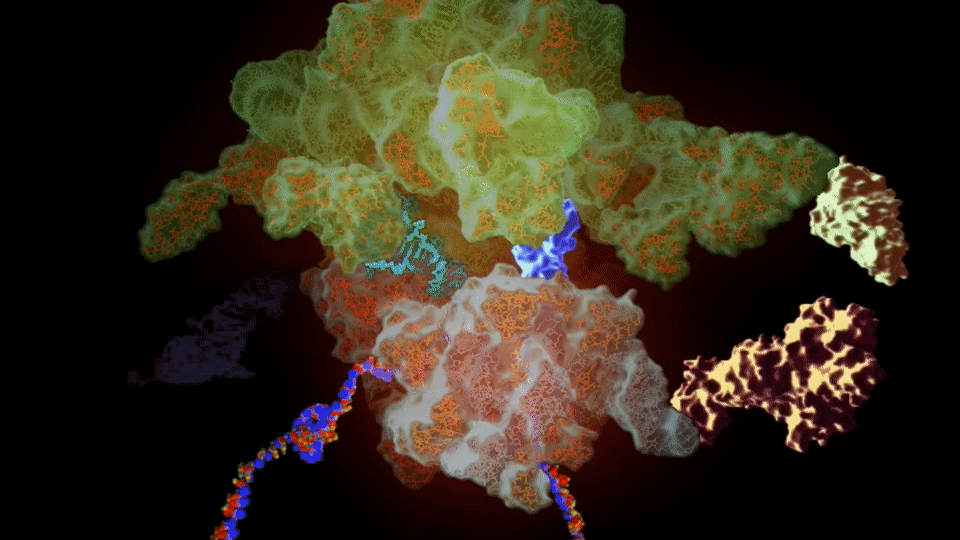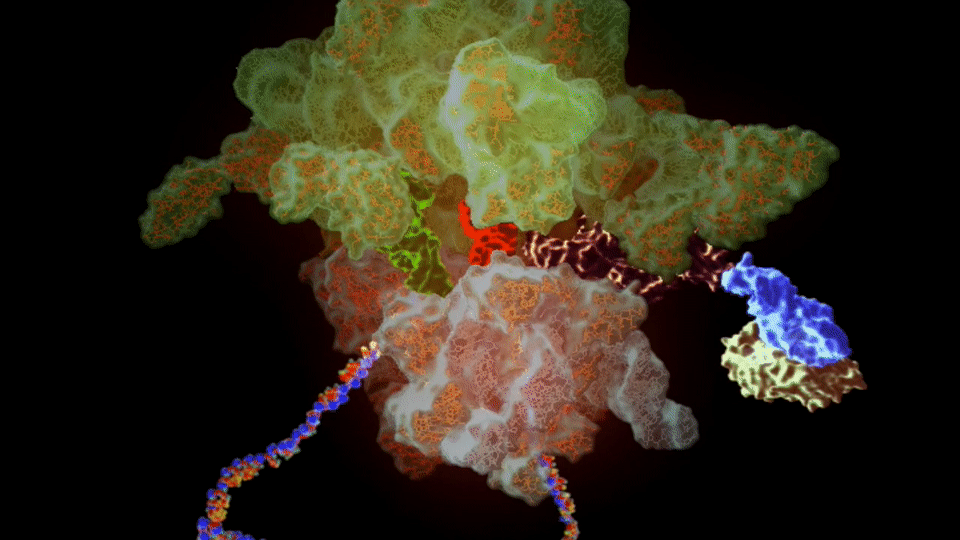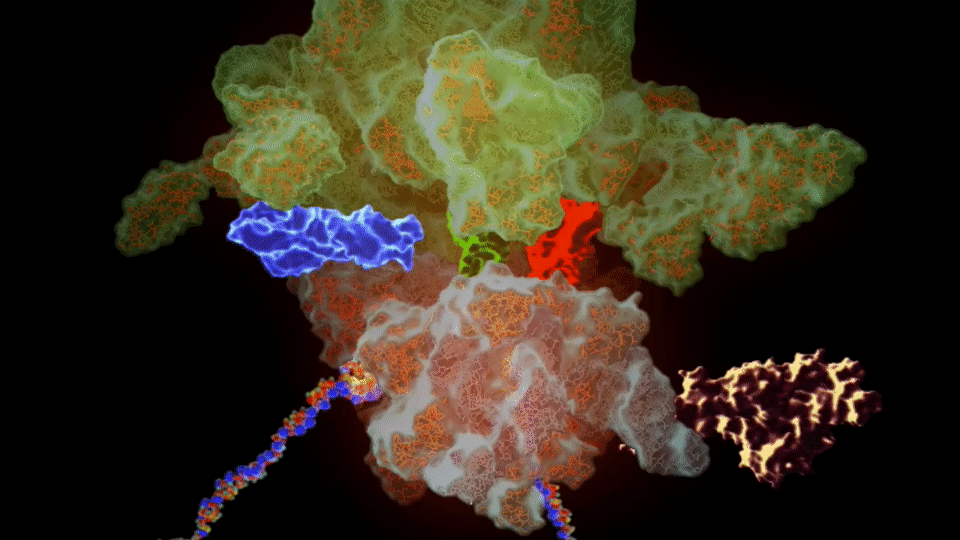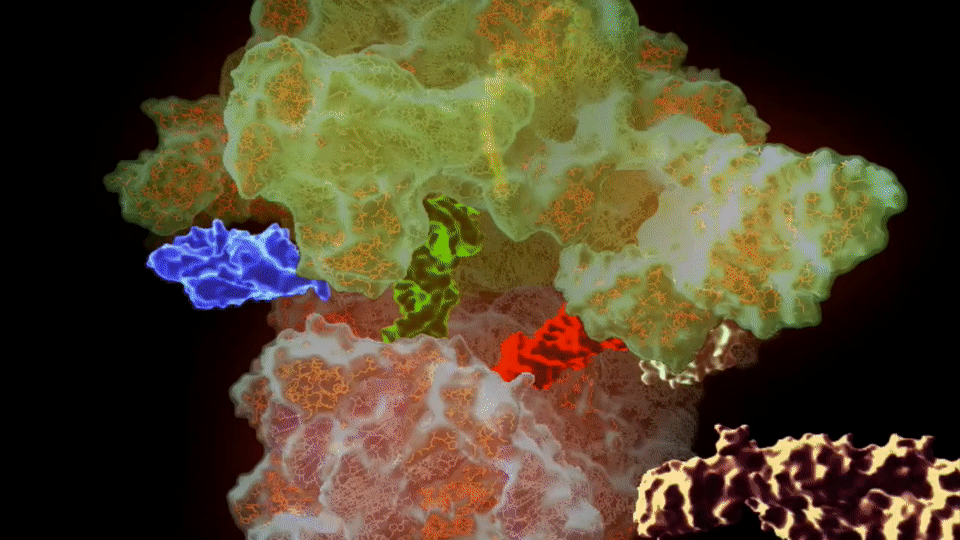
The ribosome needs various ingredients to turn genes into flesh and blood: a messenger RNA molecule, which carries the instructions for making a protein from the cellular storehouse of genetic information, held in the form of DNA. To turn this messenger RNA code into protein, the ribosome harnesses a second type of RNA – transfer RNA – which carries the building blocks of proteins, called amino acids.
Like a tape reader, the ribosome reads messenger RNA, a sequence of four “letters” (chemicals, called bases) that spells out the recipe of a protein. The chemical machinery in the ribosome takes this sequence, three letters at a time, and matches it to three letters on each transfer RNA, lining up their cargo of amino acids in the proper order, then stitches them together to build the protein.

Venki (though he has been knighted, he prefers not to be called Sir Venki) told me how his team competed to find the rhyme and reason of the ribosome against Ada Yonath, who had invested decades in cracking this problem, and Thomas Steitz of Yale University. The competition, which went on to include a fourth group in Santa Cruz, was ‘stressful’, he admitted, as it became clear that this high-risk project could blight the careers of young researchers who worked with him if it did not deliver.
They used intense, tuned sources of X-rays from a machine called a synchrotron to interrogate the structure, revealing the location of every last atom. Such studies show how the ribosome is composed of three different RNA molecules and more than 50 different proteins, arranged in two components called the 30S and the 50S subunits. The former is the ‘brain’ that reads genetic code and the latter the larger ‘heart’ that makes protein. They drift apart and together, as molecular links are forged and broken, to create the proteins that build and run the body.
Overall, the ribosome ratchets RNAs laden with amino acids through its heart at a rate of 15 per second to string them together into proteins. ‘The surprise was how a machine of this size was so precise,’ he said, while showing the Hay audience a video of this soft, deformable micro machine in action.
This is a remarkable recapitulation of the idea of the body as a machine, an ideas that was first glimpsed centuries ago, as revealed by the Science Museum’s Leonardo exhibition, and in an exhibition about humanoid robots which will open next year.
Venki told me how new understanding of this marvellous machine could help tackle superbugs that are evolving resistance to antibiotics. ‘It is potentially a huge problem and if we don’t act on it, it will become a major crisis.’
The ribosome is a big target, relatively speaking, consisting of around half a million atoms and measuring around one millionth of an inch across. The thousands at work in every one of our cells have been honed over aeons.
Studies, which date back many years, have shown which parts are best targeted by antibiotics. Peer deep inside the ribosome and, like the rings in the trunk of a tree, you can see an ancient core that has been turning instructions into the proteins for the best part of four billion years. This is ‘where all the action happens’, said Venki.
This chemical contraption lies at the heart of between two eras of life, one familiar and one shrouded in mystery: the former consists of today’s DNA-based living things and the latter the very first things that multiplied which did not rely on DNA at all.
This core is an artefact of this earlier era of evolution, when a different kind of life thrived on Earth. It consists of the genetic material RNA, a relic of the so-called ‘RNA world,’ when living things depended on this more delicate-yet-flexible kind of genetic material which not only stores information but, unlike DNA, can catalyse chemical reactions too. Finding this RNA in a ribosome is a bit like discovering a floppy disk, perhaps even a punched card, in a tablet computer.
When you compare this relic molecular machinery in myriad organisms, you make a startling discovery. ‘The closer you get to the heart of the ribosome, the more similar,’ says Venki: this ancient RNA is so essential to the process of staying alive that it is almost identical, whatever the creature.
Whether it belongs to a bacterium or a person, the RNA at the heart of ribosome ceaselessly turns genetic instructions into the protein molecules that build and run a body.
A rind of proteins evolved around this central RNA machinery to hone its performance and this differs depending on the creature – our ribosomes are almost twice as large as those of the bugs that infect us, for example. It turns out that some antibiotics work by, in effect, throwing a spanner in the works of this machine, targeting bacterial ribosomes while leaving human ribosomes intact so they do not cause side effects. ‘They bind to bacterial ribosomes, not so well to ours,’ he explained.
Thanks to this work, deeper insights are emerging that could pave the way to new classes of antibiotics. He told the Hay audience how these are desperately needed as resistance to conventional antibiotics takes hold, though he emphasized (as has the Chief Medical Officer Dame Sally Davies) that new drugs are not the whole answer – they must be accompanied by a multifaceted approach, from new diagnostics to regulating their use, not least in agriculture.
Many antibiotics such as tetracycline and streptomycin work by disrupting the conglomerates of RNA and protein in the ribosome. Venki has investigated how they do this, not least to understand how they fail. For example, some bacteria shatter antibiotic drugs into pieces (this is the cause of penicillin resistance), others add chemical groups to the ribosome, changing the target; mutate to alter the fundamental shape of the ribosome; or expel the antibiotic with molecular pumps. ‘Eventually, resistance will occur and this is simply a consequence of the fact that bacteria mutate and resistance can be selected for,’ he explained.
This provides understanding to develop new antibiotics. Tom Steitz has, for example, started a company that has developed several lead compounds that show promise. But it is a slog turning lead compounds into treatments. Venki said that drug companies have difficulty recouping the billion dollar investment when the latest antibiotics have to be reserved for the most resistant infections and, overall, are not as profitable as medications that have to be swallowed day in, day out. There should be better market mechanisms, though he added that Governments also play a key role.
The path to his Nobel Prize began with his upbringing in India, where his parents taught him that science ‘is an interesting life.’ After an unpromising start at school, notably when his parents ‘gave up on me’ between his seventh and ninth grades, a teacher helped kindle his interest in science and mathematics.
Venki went on to study for a doctorate in physics in Ohio. However, this turned out to be a false start (‘I realised that, actually, I was much more interested in biology’). Being relatively young, he decided to start again in graduate school: ‘I had to go backwards several years.’ Eventually, after gaining a doctorate in biology, he joked how he was lured from America by the ‘food, salary and weather’ to the LMB in Cambridge, where, most famously, Jim Watson and Francis Crick discovered the double helix structure of DNA.
While he had worked on parts of the ribosome in the United States, the LMB allowed him to study the molecular machine in its head-spinning entirety. ‘I had no idea how long that would take.’ The lab ‘focuses on long term challenges’ and, when it comes to its ten Nobel prizes, they ‘are simply a fringe benefit’ of its commitment to pure curiosity. At that time, Venki had not even managed to make crystals of ribosomes, a critical first step to study them with X rays. ‘I just had an idea and some hope.’
Roger Highfield interviewed Andre Geim last year at the Hay Festival about his work on graphene which also paved the way to a Nobel prize. An exhibition, Wonder Materials: Graphene and Beyond runs from July 2016 until June 2017 at the Museum of Science and Industry, Manchester.