All living things on Earth are made from protein building blocks that are created from the same 20 chemical units, called amino acids. With names like serine, leucine and alanine, nature strings together different combinations of these amino acids in proteins, whether those in hair, skin and muscle or the ones that break down food in the stomach.
Now researchers in Cambridge have found a way to create a synthetic bacterium to make proteins composed of novel amino acids, designed by scientists, that lie beyond the canonical 20 found in nature.
The synthetic bacterium was not only able to make novel kinds of protein, far more complicated than anything that can be made in the lab by chemists, but the genetic alteration rendered it virtually invincible to infection with viruses.
To carry out the feat, the team at the Medical Research Council (MRC) Laboratory of Molecular Biology led by Prof Jason Chin engineered the genetic code of a synthetic strain of the gut bacterium E. coli to include several nonstandard amino acids, akin to writing a book with several made-up letters in addition to the usual 26, from A to Z.
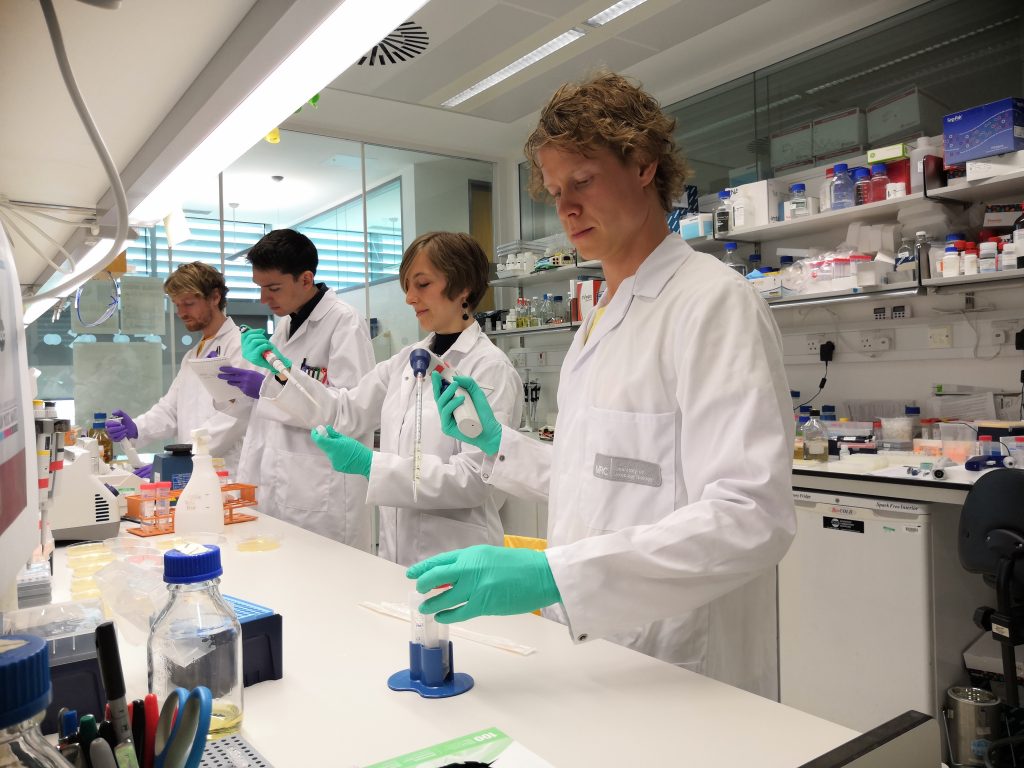

The advance rests on details of how huge molecular machines – ribosomes – that turn genetic code into protein within the bacterium read three “letters” of genetic code at a time, a “word” known as a codon, of which there are 64 different combinations.
There are only 20 amino acids because some are specified by more than one word: for example, the four different codons TCG, TCA, AGC and AGT all code for the same amino acid, serine.
In May 2019, Chin’s team reported in the journal Nature that it had simplified the E. coli’s entire genetic code, or genome, so that it packs information more efficiently into the four million letters of DNA that provide the instructions for making a cell.
His team made an organism with a radically altered genome – the largest synthetic genome ever constructed from scratch – that used three fewer codons than the usual number. Though the modified bacteria – called Syn61 – no longer had the codons in their genome they could still make normal proteins and thrive.
Now, by further engineering of their bacteria, Chin’s team has been able to use the three codons removed from the genome to encode three “noncanonical amino acids” which are not found in nature; the result is synthetic E. coli with properties not found in biology.
The researchers were able to use the synthetic bacterium to create polymers made of up to eight chemical units – monomers – strung together and join the ends to make macrocycles – a type of molecule that forms the basis of some drugs, such as certain antibiotics and cancer drugs.

Prof Chin says his research could lead to the development of novel polymers – large molecules made of many repeating units, such as proteins, plastics, and many drugs, including antibiotics.
Synthetic bacteria ‘may be turned into renewable and programmable factories that produce a wide range of new polymer molecules with novel properties, which could have benefits for biotechnology and medicine, including making new drugs, such as new antibiotics,’ he says.
‘We’d like to use these bacteria to discover and build long synthetic polymers that fold up into structures and may form new classes of materials. For example, we will investigate applications of this technology to develop novel polymers, such as biodegradable plastics, which could contribute to a circular bioeconomy.’
In the study, published yesterday in the journal Science, the Cambridge researchers also infected their synthetic bacteria with a cocktail of viruses. While natural bacteria were killed by the viruses the modified bacteria were resistant and survived.
The reason is that viruses reproduce by injecting their genetic instructions into a host cell, in this case a bacterium, and hijacking its cellular machinery.
However, when the machinery in the modified bacteria tries to read the virus genome, it fails every time it reaches the missing codons, halting the replication of the virus.
This ability to shrug off viruses is important because many drugs – for example, protein drugs, such as insulin, and polysaccharide and protein subunit vaccines, including some being developed for COVID-19 – are manufactured by growing bacteria which contain instructions to produce the drug.
‘If a virus gets into the vats of bacteria used to manufacture certain drugs, then it can destroy the whole batch,’ says Prof Chin. ‘Our modified bacterial cells could overcome this problem by being completely resistant to viruses. Because viruses use the full genetic code, the modified bacteria won’t be able to read the viral genes.’
Although some may be uneasy about where this technology is heading, Prof Chin emphasised that, since humans harnessed fire, they have realised there is an upside and a downside to innovation – it is important experiments are done in a controlled way and properly discussed and debated with the public.
The first synthetic organism was created by the American biotech entrepreneur Craig Venter, whose microbial replicant based on a copy of the genetic code of the tiny bacterium Mycoplasma genitalium was unveiled in 2010. The new work in Cambridge depends on a smaller team and is much easier to scale up, says Prof Chin.
Dr Megan Dowie, head of molecular and cellular medicine at the Medical Research Council, which funded the study to expand the lexicon of life, commented that it marks ‘a really exciting example of the value of a long-term commitment to fundamental, or discovery, science.’
Such research ‘has huge potential for major impact in biopharma and other industrial settings.’